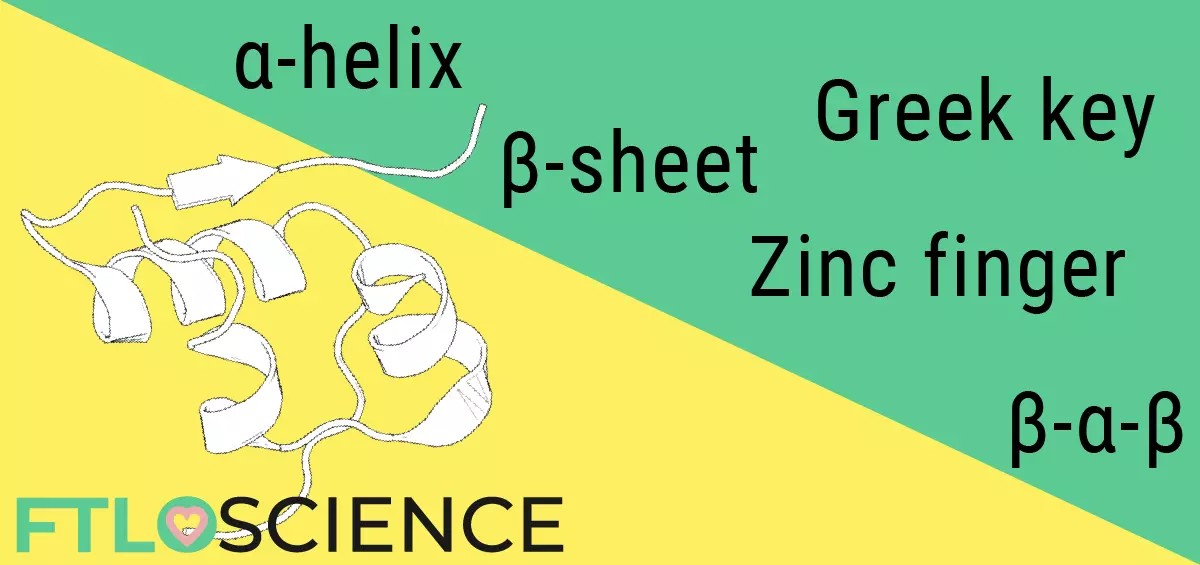
Humans have at least 20,000 different proteins scattered throughout our bodies, each with a specific role that keeps us alive and (mostly) functioning. Because of their structural diversity, proteins are often depicted as a collection of motifs like alpha helices and beta sheets, making the ability to interpret such ribbon diagrams an invaluable skill.
Amino Acids
The purpose of having genetic material—DNA—is to produce sequences of amino acids. When linked together in long chains, they become proteins, each with different structures and functions in our body. There are 20 amino acids, the ‘building blocks’ of biochemistry, some of which we can produce and some of which we need to obtain from our diet.
Each amino acid has different chemical properties, so depending on how they are linked together, they can form structurally complex and diverse proteins. Since structure correlates directly to function, biochemists need a hassle-free way to present it. Ribbon diagrams allow us to interpret the overall shape of a protein easily, without the need to account for individual amino acids.
The Protein Ribbon Diagram
If we tried to draw the chemical structure of a protein, the sheer number of atoms would make it a daunting task—insulin, considered a small protein with 51 amino acids, contains 788 atoms! Luckily, ribbon diagrams were invented by biochemist Jane Richardson in 1969, allowing us to visualize a protein’s overall structure without individual atoms getting in the way.
Below is the chemical ‘ball-and-stick’ model (left) next to the ribbon diagram (right) of an alpaca protein from The Protein Data Bank (PDB: 2XV6).
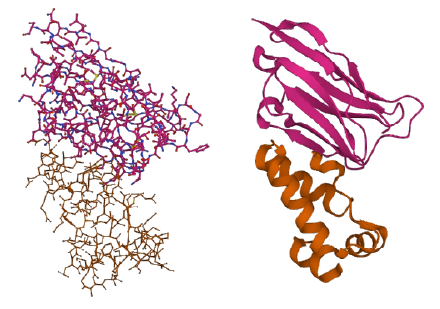
It isn’t easy to deduce the protein’s structure in the ball-and-stick model, in contrast to the ribbon diagram, which gives a much clearer view of its overall shape. That’s because the ribbon diagram only looks at the backbone of the amino acid chain and not individual atoms. We can then interpret the ribbon diagram by identifying patterns (or motifs) commonly shared among proteins.
Alpha Helices
A core component of the ribbon diagram is the alpha helix (α-helix), which forms when between 4 to 40 amino acids ‘turn’ in a helical fashion. Being stable structures, the alpha helix is a common motif found in most proteins.

Beta Sheets
Another common motif is the beta sheet (β-sheet), which is formed when multiple strands of amino acids (beta strands) bind in sequence to form a sheet structure. Each beta strand is between 3 to 10 amino acids long, with hydrogen bonds holding the entire motif together.
Hairpins and Turns
The beta-hairpin loop and helix-turn-helix are bridging motifs. The hairpin loop is the name given to amino acids that join strands that make up a beta sheet, while helix-turn-helix motifs are found between alpha helices.

Other Structural Motifs
There is no strict classification of what constitutes a motif; researchers are always looking for patterns when interpreting ribbon diagrams. Structures like Greek keys, Zinc fingers, Nests and G-quadruplexes exist. These repeating patterns often have similar functions across different proteins, such as DNA binding or fibril formation.
Software to Interpret Ribbon Diagrams
While the PDB maintains a huge protein structure resource, many different programs are available for visualizing and interpreting ribbon diagrams. Several programs include:
- Swiss-PdbViewer (view multiple proteins at the same time)
- SAMSON (with support for other non-protein structures)
- PyMOL (simple interface, beautiful renders)
- Protein Imager (browser-based)
- Nanome (Virtual Reality-enabled!)
- Mercury (access to crystal structures)
About the Author

Sean is a consultant for clients in the pharmaceutical industry and is an associate lecturer at La Trobe University, where unfortunate undergrads are subject to his ramblings on chemistry and pharmacology.